What is the Transcriptome?
In a body, a genome is everything withing chromosomes, all of the DNA the body and everything it encodes is a genome. Following transcription, DNA is processed into RNA. A body`s transcriptome is a collection of all the RNA transcribed from the genome. Genomics studies the genome and transcriptomics studies the transcriptome. This is typically done using the high throughput techniques of RNA sequencing and micro-arrays [1].
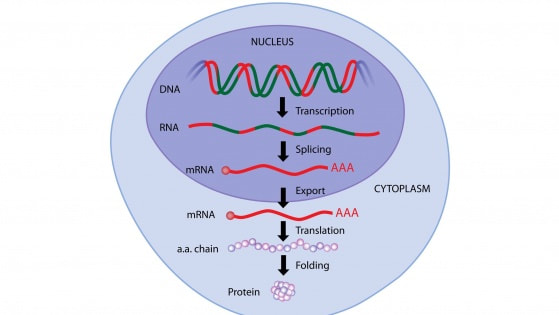
Fig. 1 This is the central dogma of biology. Transcriptomics studies RNA collectively ie. RNA, mRNA, tRNA, rRNA, etc.
How can we Study the Transcriptome?
RNA Sequencing
|
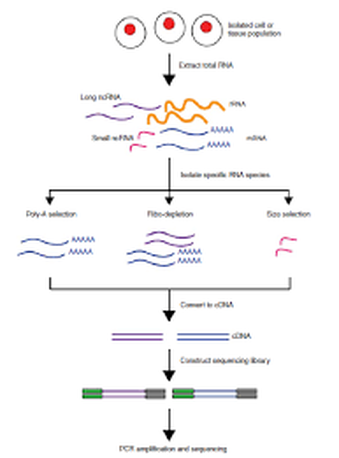
RNA-seq is a technique that can be employed to study changes in gene expression. Changes in levels of transcription correlate to differential regulation of genes. This is an incredibly powerful tool that has recently made modern transcriptomics possible. RNA-seq provides incredibly precise measurements of levels of gene transcripts, as well as their isoforms [2]. Before RNA-seq, hybridization arrays with fluorescenty labeled with a microarray have been widely used to determine transcript sequence, spliced isoforms, and to map transcribed regions to the genome. These microarrays gave high throughput results for low cost. However, in order to be performed accurately, genome sequence information was needed, and comparison of gene expression was difficult to compare. Today, RNA-seq helps to solve these problems.
|
Using RNA-seq in myocytes to study
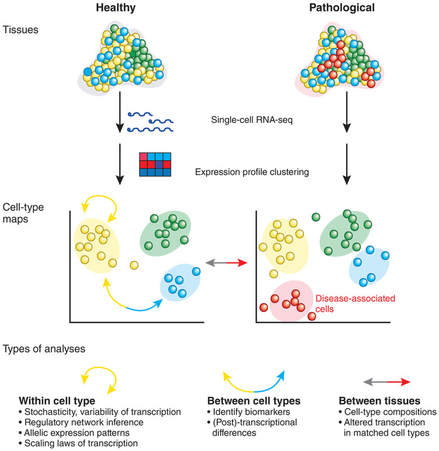
Fig. 3 Transcriptome analysis. May be used in myotubularin research to study expression between tissues to study isoform expression or expression level in muscle cells.
|
Due to work previously conducted on the myotubularin phosphatase family, MTM1 is expressed throughout the body. However, mutations in the mtm1 gene are the genetic cause of X-linked myotubular myopathy. Since the role this protein plays in cells is not completely elucidated, RNA-seq may be able to help. Since Mtm1 exists in two isoforms, it may be hypothesized that one isoform is expressed in myocytes. This isoform may contribute to healthy myocyte physiology, and since RNA-seq is sensitive enough to determine expression levels of isoforms, a connection may be made between either isoform expression or expression level in muscle cells. Alternatively, protein interactions may be skeletal muscle specific. An analysis of the transcriptome under wild type and mutated conditions may show altered gene expression. Specific protein interactors may be specific to muscle tissue.
Myotubularin-1 IsoformsThere have been 13 variants of myotubularin-1 found withing the myotubularin family. Myotubularin related proteins (MTMR1-MTMR13) all contain the GRAM domain but alter their activity as being either active or catalytically inactive. Catalytically inactive forms have been found to complex with an active form to perform the dephosphorylation reaction [3]. While myotubularin-1 is the known mutational cause of X-linked myotubular myopathy, it is interesting in that it is the only myotubularin protein that causes this debilitating disease. By further understanding the transcriptome and muscle specific gene regulation may provide answers to a treatment.
|
References:
[1] Zhang, Yue et al. “De Novo Assembly of Transcriptome and Development of Novel EST-SSR Markers in Rhododendron Rex Lévl. through Illumina Sequencing.” Frontiers in Plant Science 8 (2017): 1664. PMC. Web. 5 Apr. 2018.
[2] Wang, Zhong, Mark Gerstein, and Michael Snyder. “RNA-Seq: A Revolutionary Tool for Transcriptomics.” Nature reviews. Genetics 10.1 (2009): 57–63. PMC. Web. 5 Apr. 2018.
[3] Taylor, Gregory S., Tomohiko Maehama, and Jack E. Dixon. “Myotubularin, a Protein Tyrosine Phosphatase Mutated in Myotubular Myopathy, Dephosphorylates the Lipid Second Messenger, Phosphatidylinositol 3-Phosphate.” Proceedings of the National Academy of Sciences of the United States of America 97.16 (2000): 8910–8915. Print.
[1] Zhang, Yue et al. “De Novo Assembly of Transcriptome and Development of Novel EST-SSR Markers in Rhododendron Rex Lévl. through Illumina Sequencing.” Frontiers in Plant Science 8 (2017): 1664. PMC. Web. 5 Apr. 2018.
[2] Wang, Zhong, Mark Gerstein, and Michael Snyder. “RNA-Seq: A Revolutionary Tool for Transcriptomics.” Nature reviews. Genetics 10.1 (2009): 57–63. PMC. Web. 5 Apr. 2018.
[3] Taylor, Gregory S., Tomohiko Maehama, and Jack E. Dixon. “Myotubularin, a Protein Tyrosine Phosphatase Mutated in Myotubular Myopathy, Dephosphorylates the Lipid Second Messenger, Phosphatidylinositol 3-Phosphate.” Proceedings of the National Academy of Sciences of the United States of America 97.16 (2000): 8910–8915. Print.