What is an interaction network?
A protein interaction network provides a depiction of the connectivity between different cellular components and cell processes. Protein-protein interactions are required for nearly every process occurring in cells, and it is these interactions, that when altered, lead to many known diseases. These interaction networks help to understand cellular physiology and by further developing and characterizing these networks, poorly understood diseases may be able to be better understood and reaction pathways can be built. Once protein-protein interactions are characterized, they can be built into a protein interaction network to understand the global interactions specific proteins have among each other. Using what is known about certain protein interaction networks, patterns may be able to be seen. For example, by characterizing the interaction network of MTM1, trends in trafficking and fusion pathways can be observed and used to question how this protein is acting in muscle cell fusion, as X-linked myotubular myopathy is a congenital muscle disease.
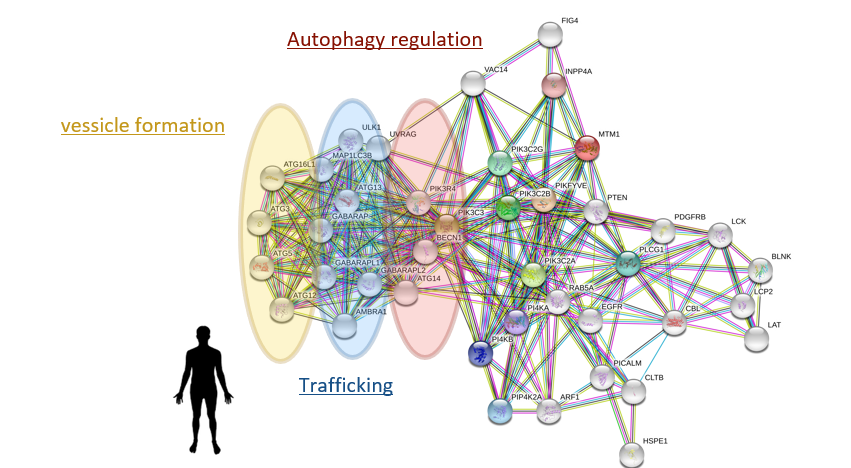
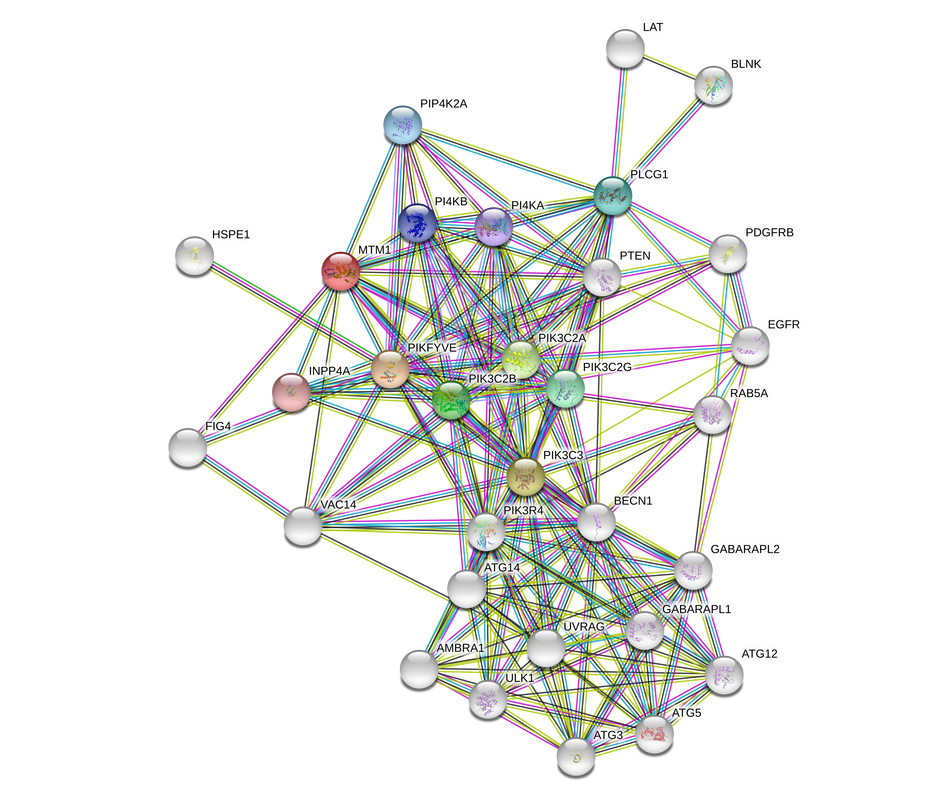
Fig. 1: An expanded view of the myotubularin-1 protein interaction network reveals proteins involved in vesicle formation and trafficking pathways. These interactions may hint at a possible role of myotubularin in muscle fusion. Patients with XLMTM show immature muscle cell fusion, and mutations in this protein could affect these pathways in a muscle cell specific manner depending on if its interactors are muscle cell specific.
How are these protein interactions found?
There are many laboratory techniques used to better characterize and understand various protein interaction networks. Some of these include protein tagging and purification, hybrid assays, and text mining.
TAP-MS |
Yeast two-hybrid screen |
|
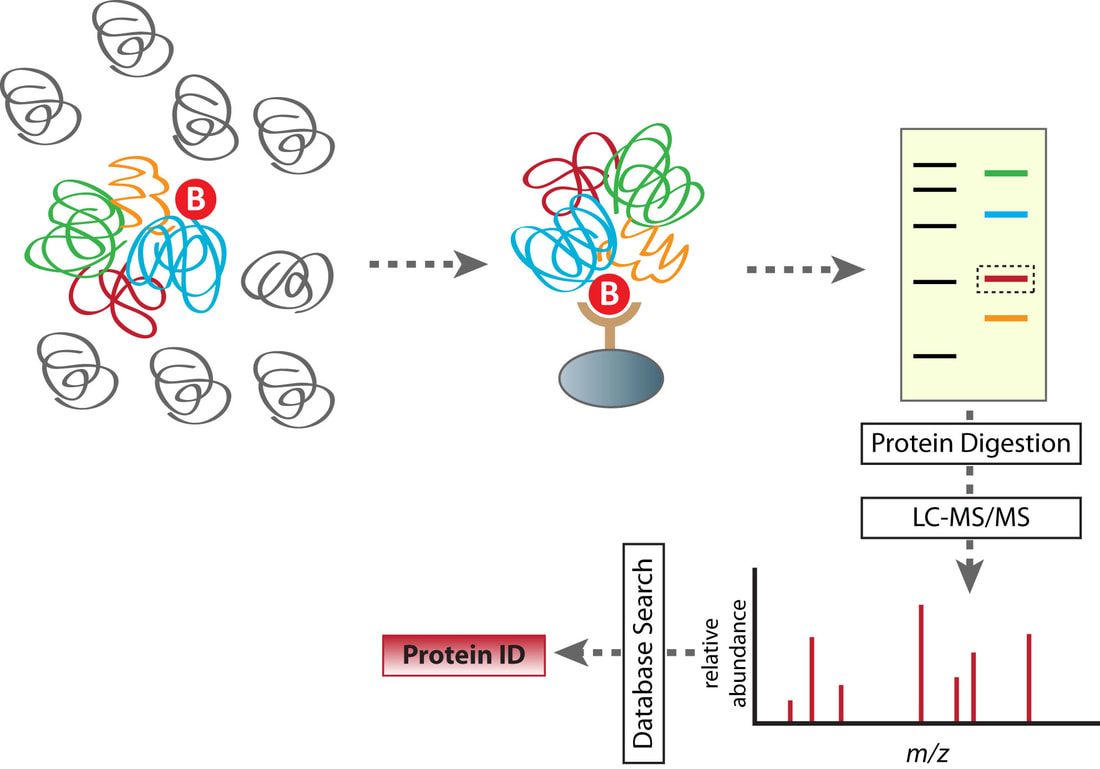
Tandem affinity purification is a method used to identify protein interaction complexes. Since protein purification methods are highly specific to the protein-protein interaction being studied, TA purification allows rapid purification of protein complexes expressed in their native state. This method relies of the fusion of a tag to a known protein on the N or C-terminal. The known tagged protein can bind to its interacting partners. The tandem part of TAP comes from two round of purification. First the interacting partners and tag are purified, then the tag is removed and the protein complex is purified from other contaminants. This method can be applied to a wide range of organisms and results in a high yield that can be broadly applied to aid in studying the proteome. Purified proteins can then be subject to mass spectrometry to identify protein interactors [2].
|
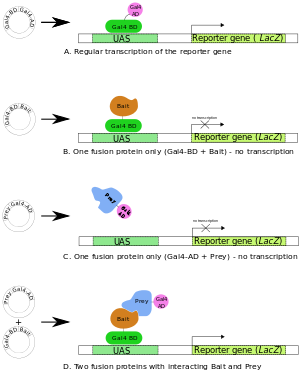
A two hybrid assay is employed to study protein-protein interactions and protein-DNA interactions. This system makes use of a protein 'bait' and 'prey' to observe coupled interactions based on the activation of a reporter in the yeast model. Identified protein interactors are them able to be purified and subject to sequencing for identification. Recently, the possibility of collecting full length protein or the 'ORFeome' is becoming a possibility. This eliminates the need for fragmentation of the protein, however, this does not cover the full proteome [1]. This is a cheap, reliable, and simple method used that is very useful for cytoplasmic protein interactions. If integral membrane proteins or membrane associated proteins are of interest, this method may give false positive results due to their cellular location and not normally being active in the cytosol.
|
Homo sapien interaction network
MTM1:
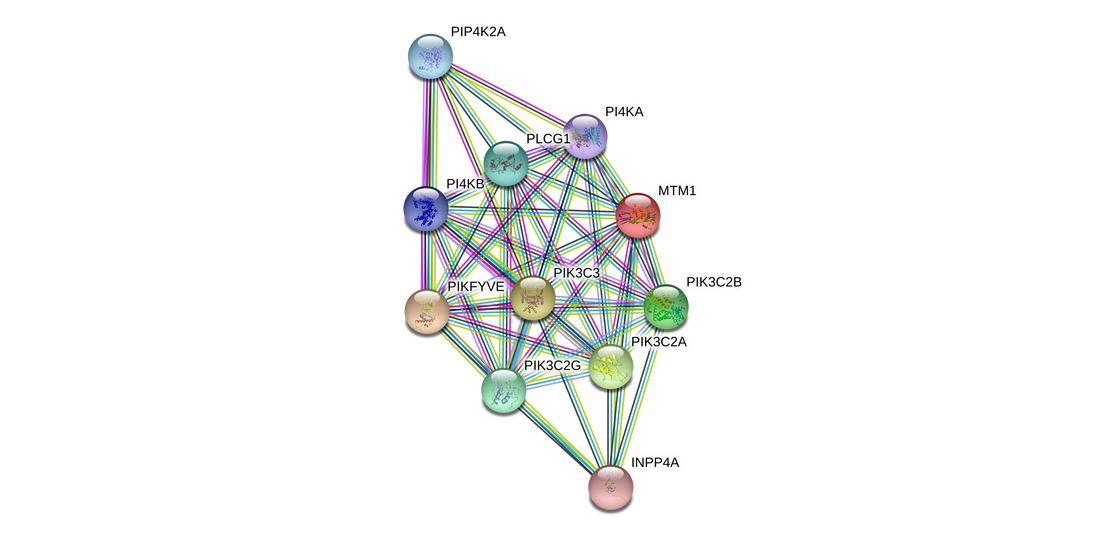
Many direct interacting partners of the human myotubularin 1 protein are kinases, as mtm1 encodes a phosphatase this makes sense. Further interaction analysis shows interaction with proteins involved in vesicle formation and trafficking.
Many direct interacting partners of the human myotubularin 1 protein are kinases, as mtm1 encodes a phosphatase this makes sense. Further interaction analysis shows interaction with proteins involved in vesicle formation and trafficking.
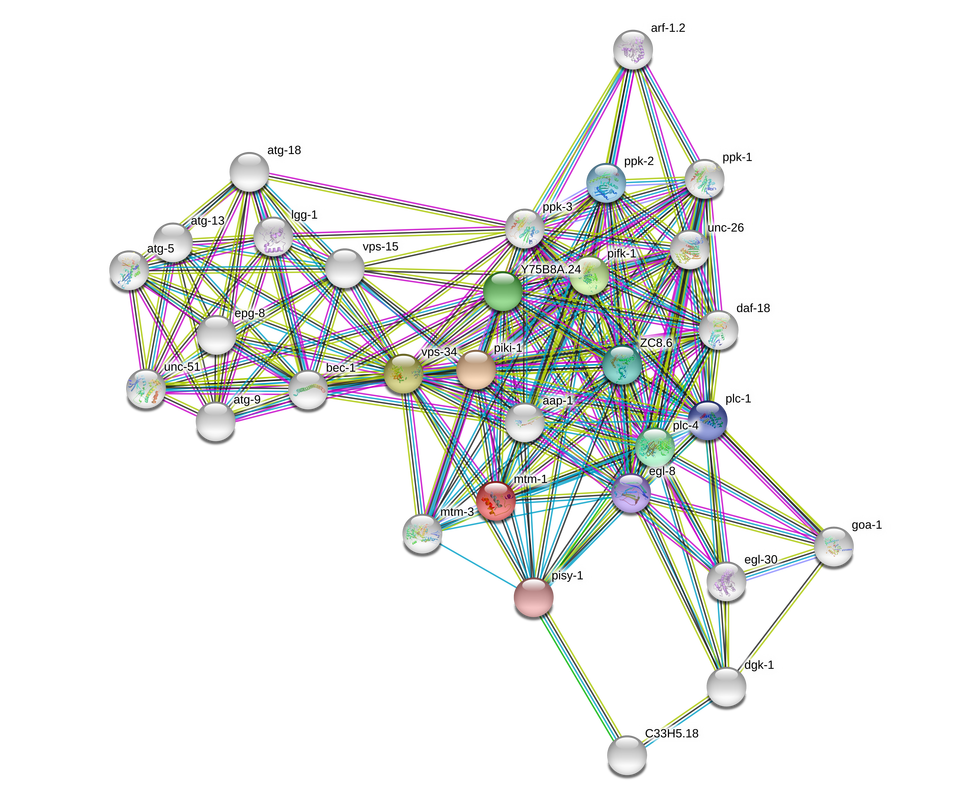
Caenorhabditis elegans interaction network
In C. elegans, the protein interaction network closely resembles that of humans but with different gene names. The proteins observed in C. elegans are initially and predominantly kinases. An expanded view shows similar proteins involved in protein trafficking and vesicle fusion. This makes C. elegans a good model to study this protein.
Conclusion:
By studying protein interaction networks, gene function and cellular role can be better understood. Myotubularin interactions with proteins involved in vesicle fusion, protein trafficking, and autophagy may hint at its role in causing X-linked myotubular myopathy. Also, by better understanding the protein interacting partners may also help understand why a mutation in this gene affects skeletal muscle development so severely when its expression in skeletal muscle is incredibly low.
By studying protein interaction networks, gene function and cellular role can be better understood. Myotubularin interactions with proteins involved in vesicle fusion, protein trafficking, and autophagy may hint at its role in causing X-linked myotubular myopathy. Also, by better understanding the protein interacting partners may also help understand why a mutation in this gene affects skeletal muscle development so severely when its expression in skeletal muscle is incredibly low.