What is Gene Ontology?
Gene ontology (GO) aims to condense the vast biological vocabulary into one database. The GO project provides researchers with ontology covering three domains; the molecular function is the gene activity at the molecular levels, the cellular component includes the extracellular environment, and the biological process of molecular events pertaining to the function of cells, tissues, and organs [1].
Molecular Function is representative of activities occurring at the molecular level such as binding or catalytic activity. Molecular functions are incredibly useful in gene understanding as these functions are typically performed by a single gene product. However, these molecular functions DO NOT where or when these functions take place, nor the context of the functions [1].
Cellular Component are the places in a cell the protein is found to act. This can also include the extracellular environment. Components may be subcellular structures or macro-molecular complexes where gene products are located. Note: this ontology includes enzymes and protein complexes but NOT individual proteins or nucleic acids. [1].
Biological Process is a series of molecular functions, defined by a beginning and end. In a broad sense, biological processes are cellular physiological processes or signal transduction. Biological processes, unlike molecular functions, MUST have more than one distinct step. Note: a biological process is NOT a pathway.
Molecular Function is representative of activities occurring at the molecular level such as binding or catalytic activity. Molecular functions are incredibly useful in gene understanding as these functions are typically performed by a single gene product. However, these molecular functions DO NOT where or when these functions take place, nor the context of the functions [1].
Cellular Component are the places in a cell the protein is found to act. This can also include the extracellular environment. Components may be subcellular structures or macro-molecular complexes where gene products are located. Note: this ontology includes enzymes and protein complexes but NOT individual proteins or nucleic acids. [1].
Biological Process is a series of molecular functions, defined by a beginning and end. In a broad sense, biological processes are cellular physiological processes or signal transduction. Biological processes, unlike molecular functions, MUST have more than one distinct step. Note: a biological process is NOT a pathway.
Molecular Function of MTM1
|
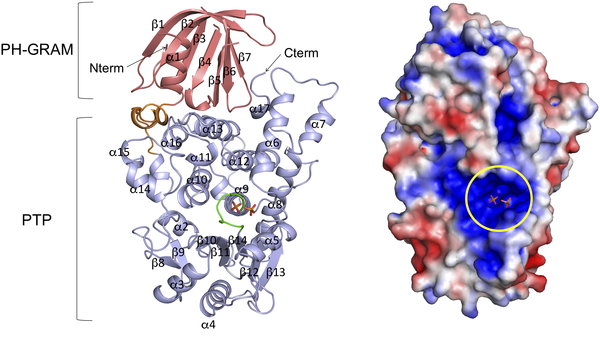
The human mtm1 gene, as well as orthologs found in dogs, cats, cows, and horses exhibit the same molecular function of human myotubularin or myotubularin related protein. These functions are phosphatidylinositol-3-phosphatase activity (GO:0004438-Canis lupus, GO:0004438-Homo sapien) mediated by the PTP phosphatase domain, and protein phosphatidylinositol binding (GO:0005515-Homo sapien, GO:0035091-Canis lupus) mediated by the PH-GRAM domain.
|
Cellular Component of MTM1
|
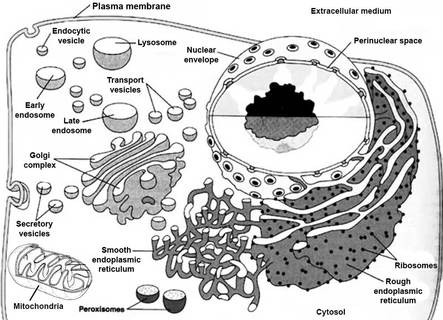
The myotubularin protein is a phosphatase located in the cytosol (GO:0005737) associating with membranes, including the phospholipid bilayer which makes up the cell membrane as well as vesicle membranes which form during endocytosis, protein trafficking, and autophagy (GO:0005770). The ontology of the cellular component is synonymous with membrane ruffle (GO:0001726).
|
Biologic Process of MTM1
|
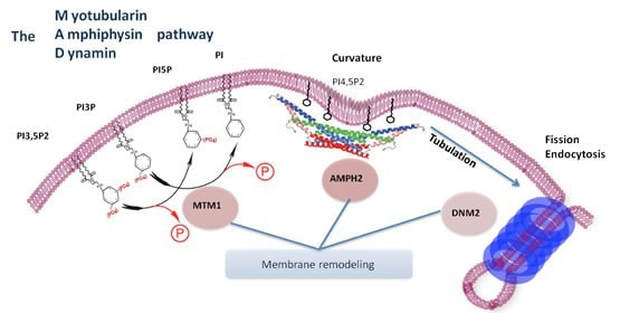
Myotubularin-1 is involved in lipid metabolism . In particular, myotubularin is a phosphatase that dephosphorylates lipid secondary messengers which is required for apoptosis, the endosomal trafficking , and autophagy [2]. As well as proper organelle morphology. MTM1 phosphatase acts in phosphoinositide dephosphorylation and phosphotidylinositol phosphate catabolism. Myotubularins phosphatase activity is the removal of a phosphate group (PO4)3− in a reversible post-translational modification. This is also involved in enzymatic activity through the detachment of phosphoric esters.
|
Conclusion:
Using Gene Ontology, various protein aspects from molecular function to biological process are condensed into one place making analysis from around the world accessible as one language. Myotubularin-1, a protein tyrosine phosphatase is involved int the metabolism of phosphatidilinositols, and in particular phophatidylinositol 3-phosphate. While functions of this protein are known, and mutations are known to cause the lethal X-linked recessive disorder X-Linked Myotubular Myopathy, the mechanisms by how this occure are undefined. GO allows us to analyse known components to hypothesize its functional mechanism. Since this protein is involved heavily in endosomal/phagosomal maturation, it may be reasonable to hypothesize a mechanism of membrane fusion.
References:
[1] ftp://ftp.geneontology.org/go/www/GO.doc.shtml
[2] The lipid phosphatase myotubularin is essential for skeletal muscle maintenance but not for myogenesis in mice Anna Buj-Bello, et al. Proceedings of the National Academy of Sciences Nov 2002, 99 (23) 15060-15065; DOI:10.1073/pnas.212498399
fig.1 Bong SM, Son K-b, Yang S-W, Park J-W, Cho J-W, Kim K-T, et al. (2016) Crystal Structure of Human Myotubularin-Related Protein 1 Provides Insight into the Structural Basis of Substrate Specificity. PLoS ONE 11(3): e0152611. https://doi.org/10.1371/journal.pone.0152611
fig.2 https://cellbiology.med.unsw.edu
fig.3 http://archive.igbmc.fr/recherche/Prog_TMN/Eq_JLapo/JL4.html
[1] ftp://ftp.geneontology.org/go/www/GO.doc.shtml
[2] The lipid phosphatase myotubularin is essential for skeletal muscle maintenance but not for myogenesis in mice Anna Buj-Bello, et al. Proceedings of the National Academy of Sciences Nov 2002, 99 (23) 15060-15065; DOI:10.1073/pnas.212498399
fig.1 Bong SM, Son K-b, Yang S-W, Park J-W, Cho J-W, Kim K-T, et al. (2016) Crystal Structure of Human Myotubularin-Related Protein 1 Provides Insight into the Structural Basis of Substrate Specificity. PLoS ONE 11(3): e0152611. https://doi.org/10.1371/journal.pone.0152611
fig.2 https://cellbiology.med.unsw.edu
fig.3 http://archive.igbmc.fr/recherche/Prog_TMN/Eq_JLapo/JL4.html